Research
Broad Research Interests of the Department
Biochemistry
Prof Adrienne L. Edkins -Cancer and Stem Cell Biology
Our group is studying the biochemistry and cell biology of the molecular chaperone, Hsp90, in cancer and stem cell biology. We use a range of biochemical (recombinant protein expression), molecular (quantitative RT-PCR, RNA interference) and cell biological (transfection, confocal microscopy and flow cytometry) techniques in order to elucidate the activity of Hsp90 and its co-chaperone, Hop, in specific cellular processes that underpin cancer and stem cell biology. In addition to our research on human chaperones, our research group is interested in comparative studies of Hsp90 complex from other organisms. These studies are interesting in that they may identify differences between the different systems that could be useful in understanding the species specific features of chaperones and may identify exploitable differences between the structure of the human and Hsp90 from other species to complement our studies to identify novel natural product inhibitors of Hsp90.
Prof Heinrich Hoppe - Malaria Parasite Cell Biology
Our group is predominantly focused on exploring ADP-ribosylation factor (ARF) GTPases as drug targets for the development of novel therapeutics against malaria and cancer. This entails the development of plate-based assays to robustly detect the activation and deactivation of ARF GTPases by regulatory proteins, implementing these assays in screens for novel ARF inhibitors and performing biochemical and cell biological experiments to elucidate the effects of the inhibitors on cultured cells/parasites. A second activity of the group is to perform routine phenotypic assays to screen compounds for activity against malaria parasites (Plasmodium falciparum), trypanosomes (Trypanosoma brucei) and mammalian cells. These screens are conducted to facilitate the vibrant drug discovery programs in the Rhodes University Centre for Chemico- and Biomedicinal Research.
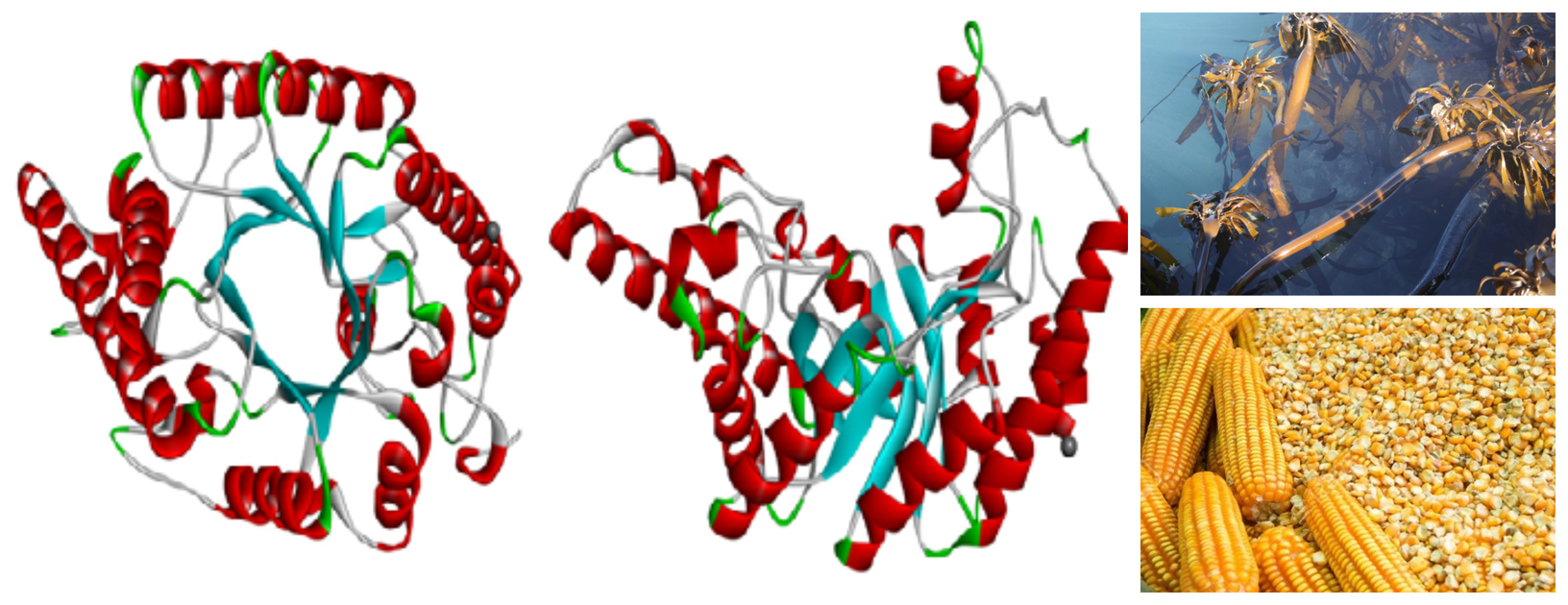
Prof Brett Pletschke - Enzyme Biotechnology, Bioproducts
Our group’s research interest lies in the field of enzyme biotechnology, more specifically in the use of enzymes in bioconversion to generate bioproducts from renewable biological resources, e.g. lignocellulose and seaweeds (macroalgae) for the bioeconomy. Since 2006 we have established ourselves firmly in the area of carbohydrate active enzymes (CAZymes) and the synergisms between these enzymes (cellulases, xylanases and mannanases) - mainly for optimising biofuel production. Currently (2019), we are looking at xylanases for improving digestibility of animal feeds, mannanases for the production of prebiotic manno-oligosaccharides (MOS), and the extraction of seaweed derived fucoidans and other enzyme inhibitors compounds from seaweeds for use as anti-diabetic/anti-obesity/anti-cancer agents.
Prof Wilhelmi's research group investigates cytochrome P450 metabolism as well as target enzymes of neurodegenerative diseases. These enzymes are cloned and expressed, purified and their activity optimised and investigated against compounds of interest. The molecular bar-coding research aims to identify specific markers in different species, including single nucleotide polymorphisms in P450, cytochrome c oxidase and other hypervariable DNA regions for both diagnostic and identification purposes.
Prof Jo-Anne de la Mare - Novel inhibitors for triple negative breast cancer
Prof de la Mare's research focuses on preclinical drug discovery for female cancers including triple negative breast cancer (TNBC) and cervical cancer. The development of drug resistance models for triple negative breast cancer and cervical cancer cell lines from South African women is funded by National Research Foundation CSUR and MRC-EC-SHIP programme grants and is geared towards developing more relevant disease models for the African context, as well as allowing for genetic characterization of the cervical cancer lines towards the identification of novel driver mutations for this disease.
Microbiology
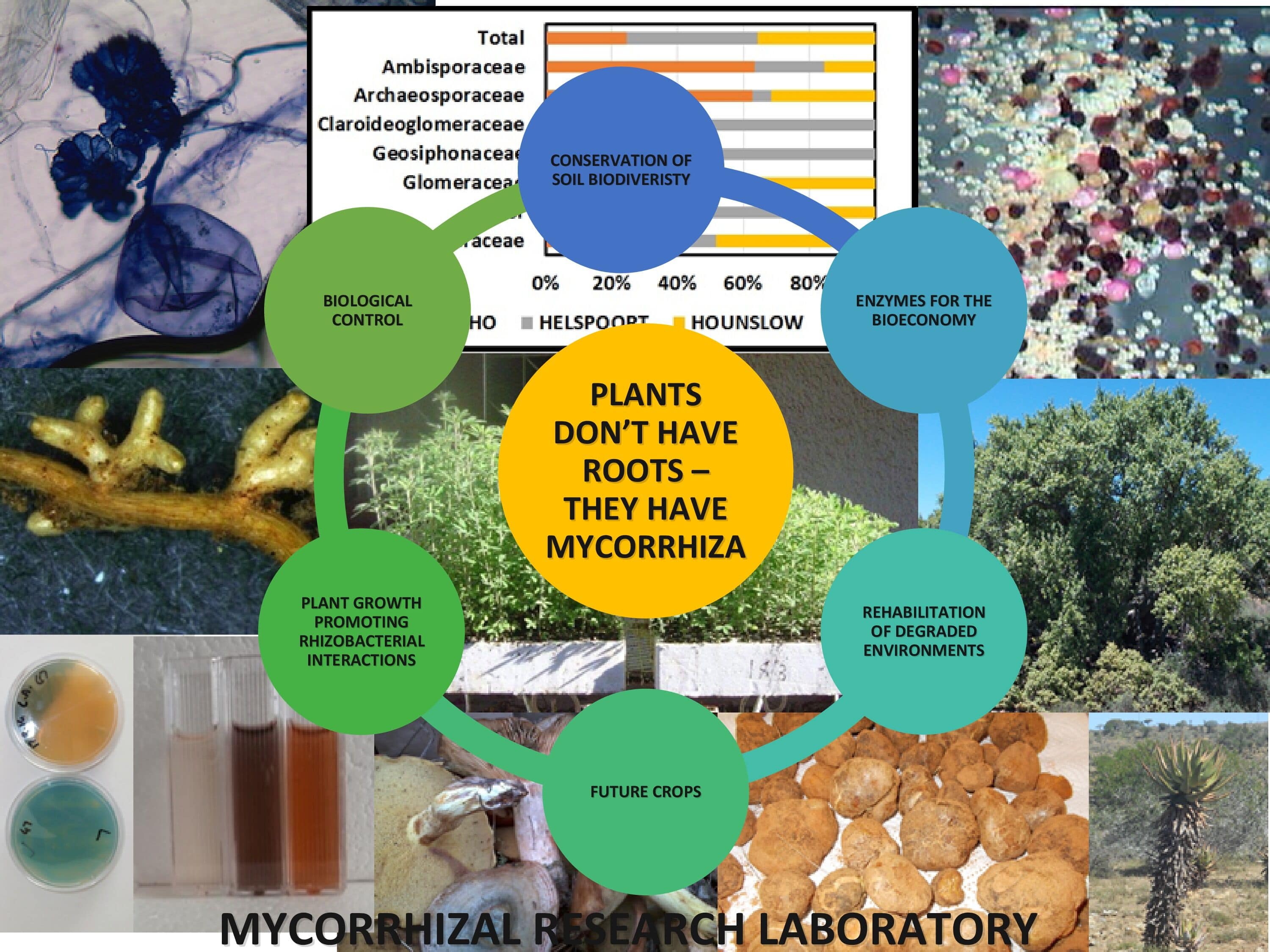
Prof Joanna Dames- Mycorrhizal Research Group
Mycorrhizal fungi are soil fungi which form a symbiotic relationship with the roots of the majority of plant species including many economically important crops. The Mycorrhizal Research Laboratory is dedicated to research in this field and research has focused on aspects of their biology, ecology, characterization, biodiversity, commercial application, their interaction with other soil microorganisms as well as benefits to both the soil environment and plant production. Mycorrhizal types of interest are the ectomycorrhizal fungi, the arbuscular mycorrhizal fungi and the ericoid mycorrhizal fungi because of their importance in the Forestry, Agriculture, Horticulture and Rehabilitation industries.
Prof Rosemary Dorrington - Microbial Ecology and Marine Natural Products Research
This research forms part of a multidisciplinary programme (Chemistry, Marine Biology Geography and Microbiology) to study the role of the microbiota (focusing on bacteria and viruses) in marine ecosystems. Research projects include the role of the microbiota in determining ecosystem health in estuarine systems, the impact of global change on marine and terrestrial ecosystems in the Southern Ocean and studies on microbial symbionts associated with marine invertebrates and their bioactive secondary metabolites.
Prof Caroline Knox - Molecular Virology
Research in Dr Knox’s laboratory is conducted in collaboration with Professor Martin Hill (Department of Zoology and Entomology, Rhodes University) and Dr Sean Moore (Citrus Research International) and focuses on the isolation and genetic characterisation of baculoviruses for the biocontrol of economically important insect pests. A second research interest, in collaboration with Professor Tom Wileman (University of East Anglia, UK) involves an investigation into the molecular mechanisms by which Picornaviruses interact with host cells during infection. Using biochemical assays and confocal microscopy, we are attempting to identify host components utilised by these viruses during the replication cycle.
One Health-related water quality and hygiene aspects, bacterial pathogenesis mechanisms, antimicrobial resistance and the transmission pathways of infectious pathogens.
Bioinformatics
Prof Ozlem Tastan Bishop - Structural bioinformatics and intelligent systems for drug-development, -resistance and -metabolism (pharmacogenomics)
A major health challenge in Africa is the underrepresentation of African populations in drug development. Pharmaceutical companies often focus on more profitable markets, neglecting diseases like malaria and tuberculosis that disproportionately affect Africans. Even when drugs exist, pathogens can develop resistance, rendering years of research and investment ineffective. Moreover, Africa’s vast genetic diversity is under-researched, and clinical trials rarely account for this, leading to potential issues with drug efficacy and safety. This is especially concerning given Africa’s high disease burden and unique genetic variation, underscoring the need for drugs tailored to African populations.
Prof. Tastan Bishop's research integrates structural bioinformatics, computational chemistry, genomics, and intelligent systems for (Afrocentric) drug development and pharmacogenomics. Her work addresses these gaps mentioned above and offers unique opportunities for South Africa, the continent, and the world. Her research aims to revolutionize drug development for diseases prevalent in Africa, pre-empt drug resistance during development, and advance pharmacogenomics by leveraging cutting-edge technologies and fostering innovation through national and international collaborations and training initiatives.
Visit RUBi webpage (click here).
Last Modified: Mon, 13 Apr 2026 15:33:40 SAST